Table Of Contents
Previous topic
Next topic
This Page
Quick search
Enter search terms or a module, class or function name.
A AtomGroup has a large number of methods attributes defined that provide information about the atoms such as names, indices, or the coordinates in the positions attribute:
>>> CA = u.selectAtoms("protein and name CA")
>>> r = CA.positions
>>> r.shape
(214, 3)
The resulting output is a numpy.ndarray. The main purpose of MDAnalysis is to get trajectory data into numpy arrays!
The coordinates positions attribute is probably the most important information that you can get from an AtomGroup.
Other quantities that can be easily calculated for a AtomGroup are
the center of mass centerOfMass() and the center of geoemtry (or centroid) centerOfGeometry() (equivalent to centroid());
the total mass totalMass();
the total charge totalCharge() (if partial charges are defined in the topology);
the radius of gyration
with radiusOfGyration();
the principal axes \(\mathbf{p}_1, \mathbf{p}_2, \mathbf{p}_1\) from principalAxes() via a diagonalization of the tensor of inertia momentOfInertia(),
where \(U\) is a rotation matrix whose columns are the eigenvectors that form the principal axes, \(\Lambda\) is the diagonal matrix of eigenvalues (sorted from largest to smallest) known as the principal moments of inertia, and \(I = \sum_{i=1}^{N} m_i [(\mathbf{r}_i\cdot\mathbf{r}_i)\sum_{\alpha=1}^3 \mathbf{e}_\alpha \otimes \mathbf{e}_\alpha - \mathbf{r}_i \otimes \mathbf{r}_i]\) is the tensor of inertia.
AdK consists of three domains:
Calculate the center of mass and the center of geometry for each of the three domains.
>>> domains = {
>>> 'CORE': u.selectAtoms("protein and (resid 1-29 or resid 60-121 or resid 160-214)"),
>>> 'NMP': u.selectAtoms("protein and resid 30-59"),
>>> 'LID': u.selectAtoms("protein and resid 122-159")
>>> }
>>> cg = dict((name, dom.centroid()) for name,dom in domains.items())
>>> cm = dict((name, dom.centerOfMass()) for name,dom in domains.items())
>>> print(cg)
{'LID': array([-15.16074944, 2.11599636, -4.37305355], dtype=float32),
'CORE': array([ 4.43884087, 2.05389476, 1.63895261], dtype=float32),
'NMP': array([ -2.99990702, -13.62531662, -2.93235731], dtype=float32)}
>>> print(cm)
{'LID': array([-15.11337499, 2.12292226, -4.40910485]),
'CORE': array([ 4.564116 , 2.08700105, 1.54992649]),
'NMP': array([ -3.20330174, -13.60247613, -3.06221538])}
What are the distances between the centers of mass?
(Hint: you can use numpy.linalg.norm() or calculate it manually.)
>>> from numpy.linalg import norm
>>> print(norm(cm['CORE'] - cm['NMP']))
18.1042626244
>>> print(norm(cm['CORE'] - cm['LID']))
20.5600339602
>>> print(norm(cm['NMP'] - cm['LID']))
19.7725089609
Does it matter to use center of mass vs center of geometry?
>>> print(norm(cg['CORE'] - cg['NMP']))
17.9463
>>> print(norm(cg['CORE'] - cg['LID']))
20.501
>>> print(norm(cg['NMP'] - cg['LID']))
19.9437
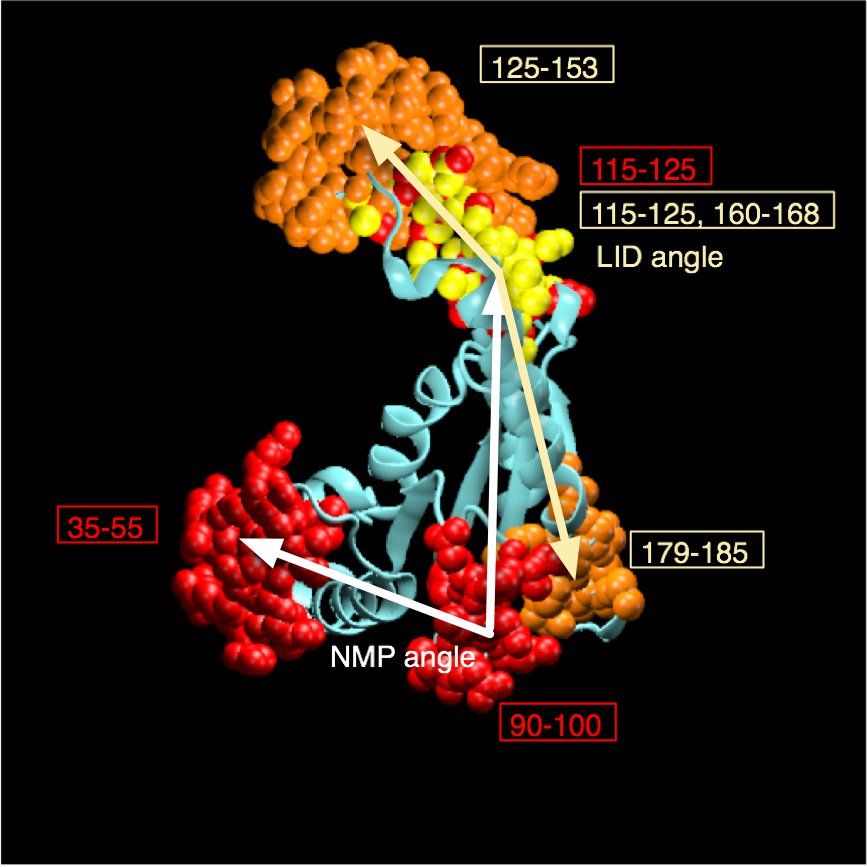
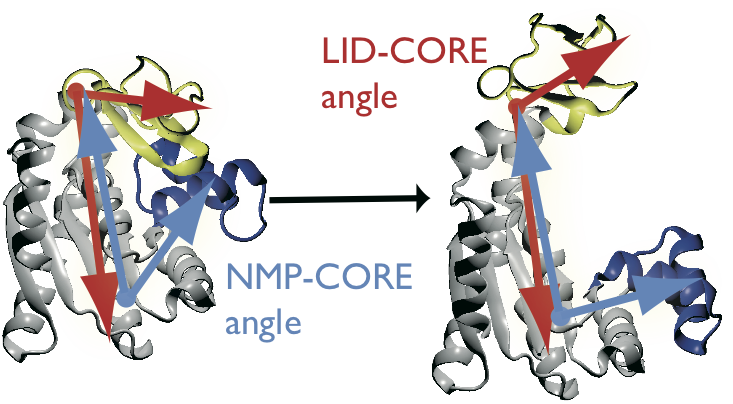
AdK undergoes a conformational transition during which the NMP and LID domain move relative to the CORE domain. The movement can be characterized by two angles, \(\theta_\text{NMP}\) and \(\theta_\text{LID}\), which are defined between the centers of geometry of the backbone and \(\text{C}_\beta\) atoms between groups of residues [Beckstein2009]:
The angle between vectors \(\vec{BA}\) and \(\vec{BC}\) is
Write a function theta_NMP() that takes a Universe as an argument and computes \(\theta_\text{NMP}\):
Use the following incomplete code as a starting point:
import numpy as np
from numpy.linalg import norm
def theta_NMP(u):
"""Calculate the NMP-CORE angle for E. coli AdK in degrees"""
A = u.selectAtoms("resid 115:125 and (backbone or name CB)").centerOfGeometry()
B =
C =
BA = A - B
BC =
theta = np.arccos(
return np.rad2deg(theta)
Write the function to file adk.py and inside ipython run the file with %run adk.py to load the function while working on it.
Test it on the AdK simulation (actually, the first frame):
>>> theta_NMP(u)
44.124821
Add the corresponding function theta_LID() to adk.py.
Test it:
>>> theta_LID(u)
107.00881
(See below for a solution.)
Calculation of the domain angles of AdK
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 | import numpy as np
from numpy.linalg import norm
def theta_NMP(u):
"""Calculate the NMP-CORE angle for E. coli AdK in degrees"""
C = u.selectAtoms("resid 115:125 and (backbone or name CB)").centerOfGeometry()
B = u.selectAtoms("resid 90:100 and (backbone or name CB)").centerOfGeometry()
A = u.selectAtoms("resid 35:55 and (backbone or name CB)").centerOfGeometry()
BA = A - B
BC = C - B
theta = np.arccos(np.dot(BA, BC)/(norm(BA)*norm(BC)))
return np.rad2deg(theta)
def theta_LID(u):
"""Calculate the LID-CORE angle for E. coli AdK in degrees"""
C = u.selectAtoms("resid 179:185 and (backbone or name CB)").centerOfGeometry()
B = u.selectAtoms("resid 115:125 and (backbone or name CB)").centerOfGeometry()
A = u.selectAtoms("resid 125:153 and (backbone or name CB)").centerOfGeometry()
BA = A - B
BC = C - B
theta = np.arccos(np.dot(BA, BC)/(norm(BA)*norm(BC)))
return np.rad2deg(theta)
|
You can directly write a AtomGroup to a file with the write() method:
CORE = u.selectAtoms("resid 1:29 or resid 60:121 or resid 160:214")
CORE.write("AdK_CORE.pdb")
(The extension determines the file type.)
You can do fairly complicated things on the fly, such as writing the hydration shell around a protein to a file
u.selectAtoms("byres (name OW and around 4.0 protein)").write("hydration_shell.pdb")
for further analysis or visualization.
You can also write Gromacs index files (in case you don’t like make_ndx...) with the write_selection() method:
CORE.write_selection("CORE.ndx", name="CORE")